All about the software for the computational biomedicine research community
The CompBioMed Software Hub addresses the needs of the computational biomedicine research community, which can use the Hub to access the resources developed, aggregated and coordinated by CompBioMed.
Core Applications
Alya
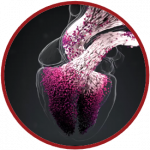
Alya, developed by the team of Mariano Vazquez and Guillaume Houzeaux at the Barcelona Supercomputing Centre, performs cardiac electro-mechanics simulations, from tissue to organ level. The simulation involves the solution of multiscale model using a FEM-based electro-mechanical coupling solver, specifically optimised for the efficient use of supercomputing resources.
HemeLB
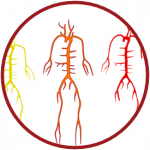
HemeLB, developed by the team of Prof. Peter Coveney at University College London (UK), is a 3D macroscopic blood flow simulation tool that has been specifically optimized to efficiently solve the large and sparse geometries characteristic of vascular geometries. It has been used to study flow in aneurysms, retinal networks, and drug delivery among many other cases.
HemoCell
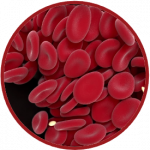
HemoCell, developed by the team of Prof Alfons Hoekstra at the University of Amsterdam (NL), is a parallel computing framework for simulation of dense deformable capsule suspensions, with special emphasis on blood flows and blood related vesicles (cells). The library implements validated mechanical models for red blood cells and can reproduce emergent transport characteristics of such complex cellular systems.
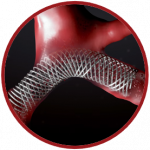
Palabos
Palabos, developed by the team of Prof Bastien Chopard at University of Geneva (CH), is a general software library for computational fluid dynamics using the lattice Boltzmann method. In biomedical research, the main domains of applications are cardiovascular, including the simulation of blood flow in arteries, the investigation of the effect of medical devices such as stents, and cell-level blood simulations to investigate fundamental blood properties.
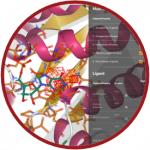
Binding Affinity Calculator
The Binding Affinity Calculator (BAC), developed by the team of Prof Peter Coveney at University College London (UK), is a workflow tool that runs and analyses simulations designed to assess how well drugs bind to their target proteins and the impact of changes to those proteins. Use of ensemble simulations to robust, accurate, and precise free energy computations from both alchemical and end-point analysis methodologies. BAC uses high-level python object abstractions for defining simulations, physical systems, and ensemble-based free energy protocols which wrap around common molecular dynamics codes.
CT2S, ARF10, BoneStrength
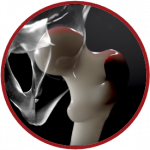
BoneStrength, CT2S, ARF10 are a set of tools to model the risk of hip fracture developed by USFD and UNIBO partners. The Computed Tomography to Strength (CT2S) workflow is a digital twin solution, used for the prediction of the risk of hip fracture for an individual based on CT scans. ARF10 is a multiscale model for the prediction of 10-years hip fracture which uses CT2S. Based on the ARF10 workflow, UNIBO is developing the in silico trial solution BoneStrength.
openBF
openBF, developed at the Insigneo Institute at the University of Sheffield (UK), is an open-source 1D blood flow solver based on MUSCL finite-volume numerical scheme, written in Julia and released under Apache 2.0 free software license. The software allows the wave propagation problem in 1D models of the human cardiovascular system to be solved, and the prediction of flow rate, pressure, and other haemodynamic variables across the network. The software has been validated against the literature in case of physiological flow in models of full body and brain circulation.
PlayMolecule
PlayMolecule is a drug discovery web service developed and maintained by project partner Acellera. It contains a variety of applications that allow users to accelerate and improve their drug discovery workflows using novel machine learning methods (such as binding affinity predictors) or through molecular dynamics simulations to elucidate biological structures and binding modes. PlayMolecule is used daily by academic institutions and industry. Both students as well as experienced researchers are using PlayMolecule to evaluate machine learning methods or simplify their molecular dynamics workflows.
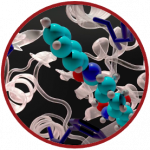
TorchMD
UPF has been focused on the continuous development of TorchMD, their software for the development and usage of machine learning potentials for molecular simulations. TorchMD is divided into three different parts/applications: TorchMD itself, which is an end-to-end differentiable molecular dynamics code; TorchMD-NET, a software for training and creating the neural network potentials; TorchMDCG, a specific application to learn coarse-grained potentials, currently specialized in protein folding.
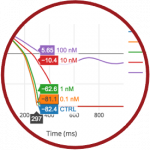
Virtual Assay
Virtual Assay is used to perform simulations of drug effects in populations of human ventricular cells, and to predict potential side effects of the drugs on the human heart. Virtual Assay has been designed to be accessible by everyone, including people with little or no expertise in computer modelling and simulations. Virtual Assay is licenced through Oxfrod University Innovation.
Associate Applications
This list includes external applications used by the consortium partners and/or developed by associate partners.